In Silico Modeling of Gastruloid Morphogenesis
Summary
One of the most fundamental events in animal development is symmetry breaking — the moment when an initially spherical ball of identical-looking cells first establishes a distinct head-to-tail axis. In gastruloids, our 3D stem cell-based embryo models, this transition is remarkably reproducible: aggregates of human embryonic stem cells reliably elongate and polarize within 48–72 hours of Wnt pathway stimulation. Yet the physical and molecular rules that make this process so consistent have remained unclear.
To investigate this, we constructed a coarse-grained agent-based computational model representing the two main cell populations found in gastruloids: peripheral (outer) cells that preferentially acquire primitive streak-like identities and high Wnt activity, and core (inner) cells that remain more pluripotent with elevated Nodal signaling. Each simulated cell is treated as a point agent interacting with its neighbors through two classes of forces: short-range contact forces governed by differential adhesion, and optional long-range chemotaxis-like forces. We then systematically swept through nearly 180,000 combinations of interaction parameters — mapping the full “morphogenetic landscape” of possible outcomes.
Differential adhesion alone is not enough
The classical explanation for cell sorting and axis formation invokes differential adhesion: cells of the same type stick more strongly to each other than to cells of a different type, causing them to segregate. Our simulations show that while differential adhesion can initiate weak asymmetries, it reliably fails to produce a single, robust axis. Even when inner and outer cells preferentially adhered to their own type, the resulting morphological asymmetry was consistently low and variable across replicate simulations. This mirrors recent experimental findings from synthetic organoid systems built purely from engineered adhesion molecules, which similarly failed to produce single, reproducible axes.
Long-range peripheral attraction is the key additional design principle
The situation changed fundamentally when we introduced a long-range attraction specifically among outer/peripheral cells — a parameter representing chemotaxis-like interactions, such as cells responding directionally to a diffusible morphogen gradient. This single additional term dramatically expanded the parameter space compatible with robust symmetry breaking: the fraction of adhesion parameter combinations yielding a well-separated, single axis grew from essentially zero to nearly 18%. The outer cells coherently converge toward one pole of the aggregate while the inner cells are displaced to the opposite pole — producing high morphological asymmetry with minimal cell loss, and doing so reliably across independent simulation replicates.
Importantly, long-range peripheral attraction does not require precise fine-tuning of adhesion: it converts a narrow, adhesion-sensitive window into a broad basin of attraction for symmetry breaking, making the outcome robust to perturbation. The optimal configurations share a consistent logic: strong inner–inner cohesion (preventing core fragmentation), weak inner–outer adhesion (allowing the two populations to separate), moderate outer–outer adhesion (permitting peripheral reorganization), and long-range outer–outer attraction that coherently pulls the periphery into a polarized architecture.
From mechanics to genetics: a minimal regulatory network
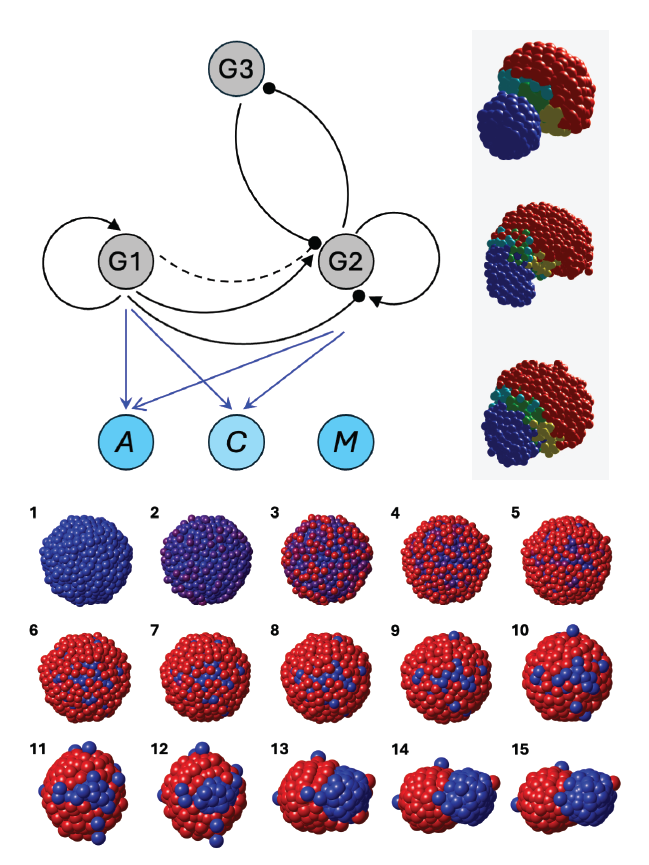
Having identified the mechanical design principles, we asked whether this behavior could arise autonomously from gene regulation — starting from a homogeneous cell population with no pre-assigned types. We designed a minimal three-gene regulatory network implementing five modular functions: (1) a timer gene with self-activation and strong degradation, creating two distinct temporal stages; (2) a diffusion-driven circuit translating the aggregate’s radial geometry into differential gene expression between inner and outer cells; (3) a bistability circuit that bifurcates two genes into distinct high/low states; (4) a lock-in circuit preserving these states after the bifurcation; and (5) a readout layer that maps gene states directly to the short- and long-range mechanical parameters.
This three-gene network successfully recapitulates the full morphogenetic sequence from scratch — a homogeneous spherical aggregate spontaneously transitions to a heterogeneous, elongated structure with a single stable axis — without any pre-specified cell types or externally imposed gradients.
DevSim: an open simulation platform
To make these modeling frameworks broadly accessible, we developed DevSim, a user-friendly MATLAB-based simulation platform for exploring arbitrary genetic-mechanical regulatory networks. DevSim allows flexible specification of population size, gene number, regulatory topology, force parameters, and noise levels, and runs on personal computers (Windows/macOS), high-performance computing clusters, and as a web browser application. A full symmetry-breaking simulation — from a homogeneous spherical aggregate to a heterogeneous elongated structure — completes in under two minutes on a standard laptop.
Representative figures



Citation
Guan G.†, Wang S.†, Shields T.G.†, Pahng S.H., Shao C.X., Ye J., Budjan C.‡, Hormoz S.‡ (2025). Cooperative short- and long-range interactions enable robust symmetry breaking and axis formation. bioRxiv. doi: 10.1101/2025.09.27.678924
† Equal contribution ‡ Co-corresponding authors