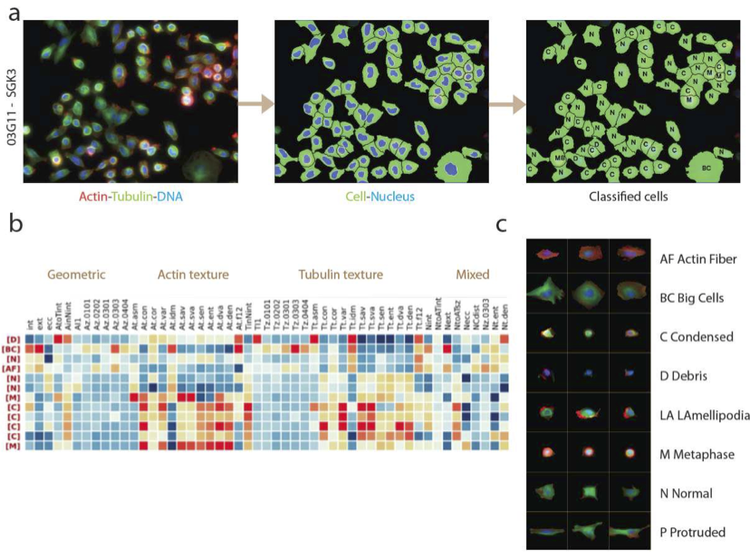
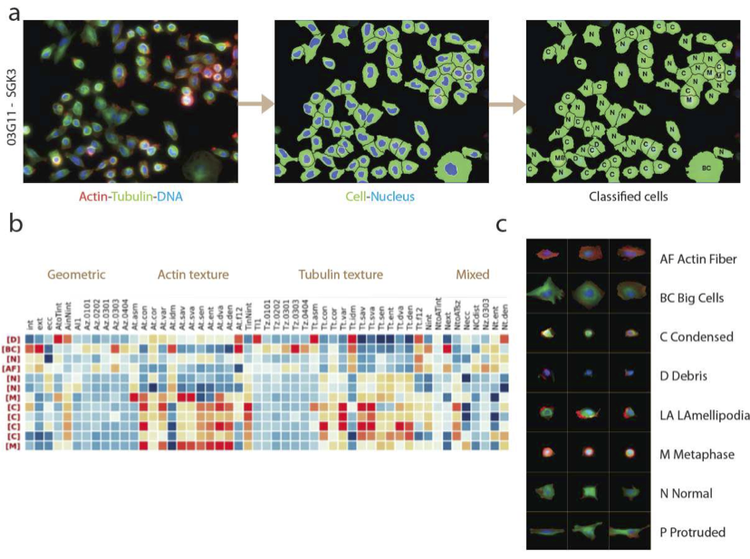
Functional genome-wide screens using RNAi and image-based phenotyping
High-throughput loss-of-function screening allows a systematic and large-scale quantitative analysis and functional annotation of all genes in the genome. With homogeneous and pathway-specific reporter assays only giving very limited information about cellular phenotypes, high-content microscopy combined with image analysis provides a multi-dimensional and quantitative single-cell readout of loss-of-function phenotypes.
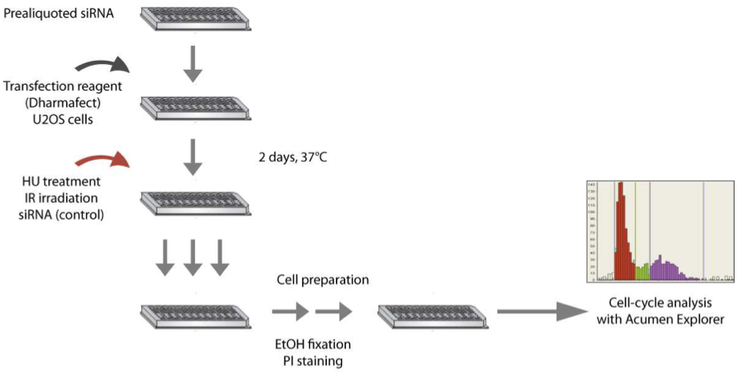
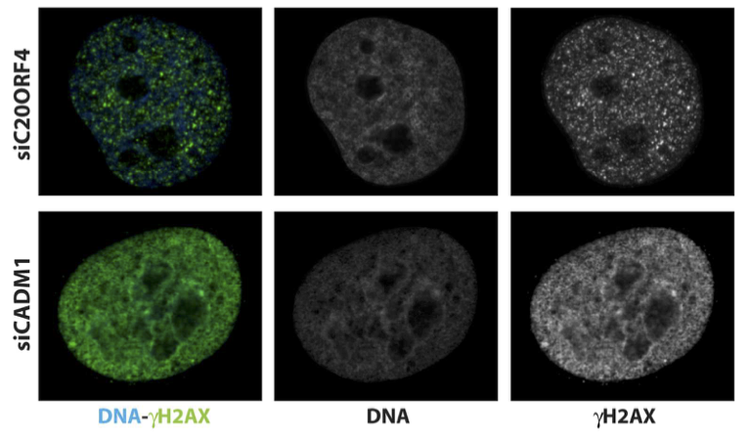
For my Bachelor thesis project I focused on functionally analyzing a set of previously unknown genes that clustered into a functional module highly enriched in genes involved in regulation of genomic integrity and the DNA damage response (DDR). Based on phenotype similarity I hypothesized that those genes might be involved in similar functions to known genes. I confirmed this hypothesis by using high-throughput in situ cytometry after knockdown of candidate genes. To compare putative conserved functions of those candidate genes I performed similar experiments using Drosophila cultured cells after depletion of homologous genes. Furthermore, because of their predicted function in DNA damage response I also checked for ectopic activation of DDR following knockdown in combination with DNA damage inducing agents.
In summary, we could show that quantitative automated analysis of perturbation phenotypes on a genome-wide scale provides an effective way of annotating unknown genes by categorizing them into known functional modules based on their phenotypic profile.
Part of the work I performed during my Bachelor thesis was later published as part of a paper by Fuchs et al. 2010 entitled Clustering phenotype populations by genome-wide RNAi and multiparametric imaging. I have also made my thesis available via figshare.
Fuchs, F.*, Pau, G.*, Kranz, D., Sklyar, O., Budjan, C., Steinbrink, S., Horn, T., Pedal, A., Huber, W. and Boutros, M. (2010). Clustering phenotype populations by genome-wide RNAi and multiparametric imaging. Mol Syst Biol 6, 370. doi: 10.1038/msb.2010.25
* Equal contribution